Tutorial
SimFluVar is freely available from the PRODUCT in SimFluVar homepage. You may download the compressed SimFluVar files for your operating systems. Detailed installation method for each operating system is as follows.
More Detail
1. Program Introduction Figure1. Screen a running process of SimFluVar programe
Figure2. Computational procedure of gene modification patterns using SimFluVar programe
SimFluVar is a program in the SimFlu (Simulation tool for influenza virus) series; its modification parameters, in 61x61 formations, are calculated based on year-to-year modification patterns extracted from empirically discovered influenza virus gene information.
This programs output is codon-level computation of the modification patterns with time on influenza A virus classified by year; the program also provides analysis results on change in base creation within wobble base position of each codon.

SimFluVar program can be utilized for computer simulation or to analyze gene modification patterns that exist among time-series influenza virus outbreak groups. Detailed gene modification computation process is as follows.
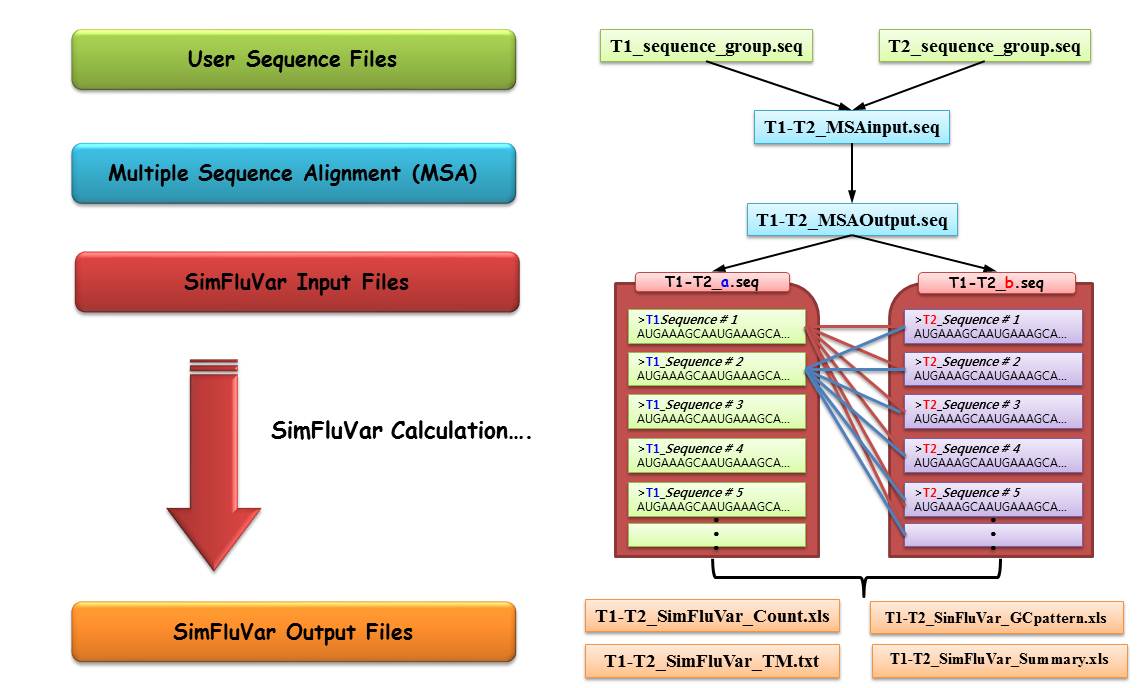
First, to activate SimFluVar program, 'multiple sequence alignment' (MSA) needs to be executed between sequences from 2 subject years. For the example illustrated above, sequences for time 'T1' and time 'T2' were named 'T1_sequence_group.seq' and 'T2_sequence_group.seq', respectively.
Following the above steps, and if the MSA process for T1 and T2 groups has been successfully executed, it can be construed that the preparation required for activating SimFluVar program is completed. To start SimFluVar, the result file containing gap generated from MSA needs to be divided into two separate files, each containing individual time, T1 and T2; it should be noted that as SimFluVar automatically searches and registers files with '*.seq' extension in the [user_input] folder within the user-designated folder, the user must create 2 files using the file extension. For the above example, in which the file was separated into 'T1-T2_a.seq' and 'T1-T2_b.seq', the suffix 'a' and 'b' before the file extension should be noted. When searching input files, SimFluVar program looks for *.seq format files; it then recognizes files with the same file name before the extension as a pair. In other words, the two files returned from searching a file name 'T1-T2_' become a pair, and the initial outbreak year and the final outbreak year are distinguished by this suffix; the suffix 'a' represents the initial time group (time T1 in the example), while the suffix 'b' represents the final time group (time T2 in the example).
First, once the input pair gets recognized and a sequential context is established, SimFluVar grasps modification patterns of each codon location by comparing all sequences of T1 time slot with all sequences of T2 time slot at the corresponding codon level. In other words, as illustrated in the above diagram, the sequences of T1 time slot (T1_Sequence #1, #2, …, #n, where n represents total the number of sequences in T1 time slot) and the sequences of T2 time slot (T2_Sequence #1, #2, …, #p, where p represents total the number of sequences in T2 time slot) make up a Counting Matrix by measuring the modifications of codons at each codon location; at this time, as the sequences of T1 time slot and those of T2 time slot are individually compared, resulting in n x p instances of comparison. Subsequently, Counting Matrix generated from outbreaks in consecutive years is converted to Markov model's Transition Matrix (TM) according to the following formula.
